 Home
Home
Shoot for the moon. Even if you miss it,
you will land among the stars.
Les Brown
Day 7: DNA assembly and DNA design...in progress...
Assignments Due
Overview of Experiment
Today you use electroporation to transform bacteria with
mutated plasmids, learn how to use a fluorescent plate reader
to measure fluorescence of purified GFP and the output of pTetR-GFP
(transformation with BioBrick).
YOU WILL HAVE TO
COME IN THE DAY BEFORE LAB DAY 7 AND PULL COLONIES:
BBa_I13522 (pTetR-GFP):
pick FOUR colonies per team and grow overnight in 3 ml LB-Amp
(50 µg/ml) at 37°C with shaking (250
rpm)
- With your Sharpie, circle well-isolated colonies on the
plates
- Put 3 ml of LB + antibiotic in sterile plastic tubes (label
as needed directly on the tubes)
- Gently swab a circled colony with a sterile pipet tip
or toothpick--do not touch any other colonies
- Eject the tip into the appropriately labeled tube
- Slightly loosen the lids and put the tubes in the 37°C
shaker overnight
- Store the plates at 4°C.
Electroporation of Mutated Plasmids
- Prechill two 1 mm electroporation cuvettes and microcentrifuge
tubes on ice
- Thaw the electroporation-competent cells (NEB
5-alpha Electrocompetent E. coli)
on ice (~10 min.) and mix by gently flicking
 You must
be extremely gentle when working with competent cells.
These cells are highly sensitive to temperature changes
and/or mechanical lysis. Mix cells by gently tapping the
tube or swirling with a pipet tip, not by pipetting up & down
or vortexing.
You must
be extremely gentle when working with competent cells.
These cells are highly sensitive to temperature changes
and/or mechanical lysis. Mix cells by gently tapping the
tube or swirling with a pipet tip, not by pipetting up & down
or vortexing.
- Transfer 45 µl of cells to a chilled microcentrifuge
tube
- Add 1 µl of PCR reaction to cells and gently mix
- Place the chilled cuvette on its side and carefully transfer
cells + DNA without
introducing bubbles
- Gently tap the cuvette until the mixture of cells and DNA settles evenly to the bottom (i.e., there is no gap across the cuvette)
- Wipe outside of cuvette with KimWipe and slide the cuvette into the electroporation chamber until the cuvette connects with the electrical contacts
- Pulse sample ONCE at 1.7 kV, 200 ohms, 25 µF
(Bio-Rad Electroporator); typical time constant = 4.8-5.1
milliseconds
**Record the time constant (in ms) and the actual volts (kV) delivered to sample**
 time constant (τ): the amount of time required for the actual voltage of the delivered pulse to decay to 1/e (37%) of the initial voltage
{τ = Resistance (R) x Capacitance (C)}
time constant (τ): the amount of time required for the actual voltage of the delivered pulse to decay to 1/e (37%) of the initial voltage
{τ = Resistance (R) x Capacitance (C)}
- QUICKLY remove the cuvette from the chamber and add 950 µl
of prewarmed (40-44°C) sterile SOC medium to the cells
- With a sterile Pasteur pipet, quickly but gently resuspend
the cells and transfer the cell suspension to a 17 mm x 100
mm round-bottom culture tube
- Incubate the sample at 37°C for 1 hour with shaking at
250 rpm
- Pipet 200 µl of transformed cells on a LB-ampicillin
plate
- Pour ~10 sterile solid glass beads onto the plate, set the
plate on the benchtop, and "shake" plate in a perpendicular
motion; invert plate to pour off beads (collect in a large
beaker -- these can be reused)
- Pellet the cells for 20 sec at 5,000xg, remove the supernatant,
and resuspend in 200 µl SOC
- On a second LB-amp plate, pipet 200 µl of resuspended
cells and REPEAT step 13.
- Let the plates sit 5 minutes at room temperature so that the liquid absorbs into the agar
- Incubate the plates (upside down) overnight at 37°C
Fluorescent plate reader training (Keck 201)
A. Fluorescence of purified GFP
Make 10-fold and 100-fold, dilutions of the stock
GFP in PBS. In a 96-well plate, prepare an array of
dilutions of purified GFP:
- plate 100 µl of each dilution in triplicate
- plate 100 µl PBS in triplicate as a control
B. Functional analysis of simple circuit: BBa_I13522 (pTetR-GFP)
Experimental goal: Does the genetic circuit work?
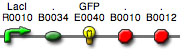
You used whole plasmid PCR to mutate the RBS for GFP in BBa_I13522 (pTetR-GFP).
This BioBrick contains BBa_E0840,
which has the coding sequence for BBa_E0040:
GFPmut3b (derived from
the jellyfish Aequeora victoria). Read the information
about BBa_E0040 at the Registry of Standard
Biological Parts: the GFPaav variant (Appl. Environ. Microbio.
(1998) 64:2240) has a half-life of around 90 minutes; GFPmut3b
has a reported excitation maximum of 501 nm and an emission maximum
of 511 nm. Today, you wil measure GFP output of the unmutated
RBS.
- Outline of goals: How do you get a big signal?
- How do you know that what you're measuring is actually
what you want?
- What are the appropriate controls?
- How do we account for background?
- How does signal noise affect the results?
- Fluorescence signal detection
- wash vs. unwashed cells
- determine density of cultures
- determine fluorescence output
Prepare cells: cells + R0040 (promoter only); cells +
I13522 (GFP)
- In a 1.5 ml tube, pellet 500 µl of each culture at
5,000xg for 20 sec
- Carefully remove supernatant and resuspend pellet in 500 µl PBS or 25% glycerol
- Prepare a 1:10 sample in a second 1.5 ml tube by adding 100 µl resuspended cells to 900 µl PBS or glycerol
- Plate 100 µl of resuspended cells in triplicate in a 96-well plate (one set for each culture and one set for each 1:10 culture)
- CONTROLS: plate 100 µl
of unwashed cells in triplicate (one set for each culture)
and 100 µl
LB in triplicate
C. Measure fluorescence
- Measure absorbance at 600 nm for all wells
- Perform a single excitation at 480 nm and measure emission
at 510 nm
- Perform an emission scan: hit at 480 nm and measure
from 500-580 nm
- Perform an excitation scan: excite from 400-489 nm and
measure at 510 nm
D. Analyze data
- Normalize for cell density
- Did you observe a signal above background?
- Look at the emission and excitation scans for the cells:
do they look like purified GFP?
- Plot excitation and emission scans on same graph
- Calculate GFP spectrum for cells + I13522 (subtract spectrum
for cells + R0040)
Copyright, Acknowledgements,
and Intended Use
Created by B. Beason (bbeason@rice.edu),
Rice University, 21 November 2007
Updated 13 November 2013

 Home
Home