 Home
Home
It's kind of fun to do the impossible.
Walt Disney
BIOC 313 Introductory Synthetic Biology
Instructor:
- Beth Beason: bbeason@rice.edu; 713-348-2535; Anderson
Biological Laboratories (ABL) 326
Time and Location: Classes meet
for 4 weeks on Tuesday and Thursday in the 2nd half
of fall semester from 1 - 5 p.m. in Biology Basement
Teaching Labs; lab begins the week after midterm recess.
Prerequisites:
BIOS/BIOC 211: Experimental Biosciences or permission
of instructor.
Registration: You may register on Esther. Enrollment
is limited to 24 students.
General Course Description: This course is intended to introduce students to the emerging field of synthetic biology. Students will present current literature that focuses on genetic parts that are currently used to program bacteria (sensors, logic functions, and actuators) and bacteria that have been successfully programmed to exhibit novel functions. The laboratory will expose students to molecular biological procedures that are routinely used in building and characterizing synthetic genetic circuits.
Preparation:
In preparation for lectures and student presentations,
everyone is expected to read appropriate background material
(Ptashne text and review articles) and the assigned paper(s) (available
in OWL-Space Resources). See How
to Read a Scientific Article (in OWL-Space Resources) for tips on reading
research papers. Additionally, you
must come to lab prepared--this requires
you to READ the
experimental protocols on the course web site BEFORE coming
to lab, not just print a copy of them and bring it with you.
Bring only the information you need to perform the experiments.
The procedures for each day are available from the Course Schedule
page, and you will be given any additional information in the
pre-lab lectures.
AIMS of Lab:
I) DNA manipulation, verification, and assembly (purification
of plasmid DNA, DNA fragmentation, size analysis of DNA fragments,
classic assembly of DNA fragments)
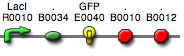
Build a simple genetic circuit using
the following BioBricks and BioBrick
Standard Assembly:
BBa_R0040 promoter/tetR,
negative (part = 54 bp) [in pSB1A2, AmpR]

BBa_E0840 GFP
reporter with strong RBS (part = 878 bp) [in pSB1A2, AmpR]
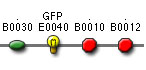
- Day 1: isolate plasmid DNA, digest BioBricks,
dephosphorylate vector
- Day 2: analyze DNA fragments on agarose
gel, purify DNA fragments from gel, ligate insert and vector,
transform bacteria with ligated DNA
- Day 3: PCR screen bacterial colonies
for those with desired inserts
- Day 4: analyze PCR products on agarose
gel
II) DNA design
(primer design, PCR synthesis of plasmids with mutated RBS,
transformation with mutated plasmids)
- Day 3: design PCR primers to mutate
RBS
- Afternoon before Day 4: pick colonies
for overnight cultures
- Day 4: isolate plasmid DNA from transformed
colony
- Day 5: perform PCR of plasmid DNA to
mutate RBS
- Day 6:
analyze PCR products on agarose
gel, digest with DpnI, clean-up DNA
- Day 7: transform bacteria with mutated
plasmid
III) Functional anaylsis of simple circuit
(fluorescence output compared with a control; characterization
of functional properties)
- Day 6: design experiment to determine
effect of mutations in the ribosomal binding site (RBS)
on GFP production
- Afternoon before Day 7: pick colonies
for overnight cultures
- Day 7: learn how to run fluorescent
plate reader (purified GFP); harvest cells, analyze fluorescence
of unwashed and washed cells
- Afternoon before Day 8: pick colonies
for overnight cultures
- Day 8: characterize fluorescence of cells
with mutated RBS
Journal Club
Presentation: Each team will
choose scientific articles in synthetic biology
to present; you and a partner(s) will give a 15 minute
PowerPoint presentation
Project Proposals:
Undergraduate
teams are charged with proposing an idea that promotes
the development of novel biotechnology.
These projects should focus on using
standardized biological parts and
simple mathematical models to design and optimize
novel genetic circuits. Although additional biological
parts can be used in these projects, the core idea
of the project cannot depend solely on the function
of these non-standardized parts. Each TEAM will prepare
a document
that summarizes the idea and contains a clear justification
for building this circuit (in the context of previously
published work), an outline of the biological parts
required for the project, and a description of the
models that will be required to build the circuit.
Assignments & Grading: Your
final grade will be determined from your lab notebook,
homework assignment(s), lab performance, Journal Club
Presentation, class participation, and Project Presentation
and Proposal.
We would like to thank New England Biolabs for their generous support of this laboratory course

Copyright, Acknowledgements,
and Intended Use
Created by B. Beason (bbeason@rice.edu),
Rice University, 10 January 2008
Updated 19 October 2013

 Home
Home